# ======================== R版ATT-A-IPTW完整分析(最终稳定版) ========================
# 1. 安装并加载所有依赖包(显式加载grid包,修复textGrob缺失)
install.packages(c("readxl", "writexl", "dplyr", "tidyr", "ggplot2", "gridExtra",
"Matching", "WeightIt", "cobalt", "mice", "boot", "broom",
"moments", "pROC", "ggpubr", "kableExtra", "grid"), # 新增grid包
quiet = TRUE, dependencies = TRUE)
library(readxl)
library(writexl)
library(dplyr)
library(tidyr)
library(ggplot2)
library(gridExtra)
library(grid) # 显式加载grid包(textGrob/gpar依赖)
library(Matching)
library(WeightIt)
library(cobalt)
library(mice)
library(boot)
library(broom)
library(moments)
library(pROC)
library(ggpubr)
library(kableExtra)
# 2. 全局环境配置(适配所有R环境)
Sys.setlocale("LC_ALL", "zh_CN.UTF-8") # 中文显示(如报错可注释此行)
options(repr.plot.width=18, repr.plot.height=10) # 绘图尺寸
options(warn=-1) # 关闭无关警告
# 基础配色方案(无需viridis包)
colors <- list(
外科 = "#E74C3C", 内镜 = "#3498DB", 截断前 = "#F39C12",
截断后 = "#27AE60", 参考线 = "#95A5A6"
)
# 3. 路径与参数配置(请修改为你的实际路径)
DATA_PATH <- "/Users/wangguotao/Downloads/ISAR/Doctor/数据分析总表.xlsx"
OUTPUT_DIR <- "/Users/wangguotao/Downloads/ISAR/Doctor/ATT_IPTW_融合分析结果_R/"
if (!dir.exists(OUTPUT_DIR)) {
dir.create(OUTPUT_DIR, recursive = TRUE, showWarnings = FALSE)
cat(sprintf("✅ 自动创建结果目录:%s\n", OUTPUT_DIR))
}
FIXED_COVARIATES <- c("lesion_max_diameter_mm", "modified_ctsi_score",
"lesion_location", "bmi", "comorbidity_hypertension", "walled_necrosis")
# 4. 函数1:列名模糊匹配
match_column_names <- function(df_raw) {
cat("\n📊 原始Excel文件列名(共", ncol(df_raw), "个):\n")
for (i in 1:ncol(df_raw)) {
cat(sprintf(" %2d. %s\n", i, colnames(df_raw)[i]))
}
# 匹配结局变量:影像学缓解
imaging_cols <- grep("影像.*缓解|缓解.*影像", colnames(df_raw), value = TRUE)
if (length(imaging_cols) == 0) {
cat("\n⚠️ 未自动匹配到'影像学缓解'列,请输入列名序号:")
selected_idx <- as.integer(readline(prompt = " 序号:"))
imaging_col <- colnames(df_raw)[selected_idx]
} else {
imaging_col <- imaging_cols[1]
cat(sprintf("\n✅ 自动匹配'影像学缓解'列:%s → imaging_response\n", imaging_col))
}
# 匹配处理变量:手术方式
treatment_cols <- grep("手术.*方式|方式.*手术", colnames(df_raw), value = TRUE)
if (length(treatment_cols) == 0) {
cat("\n⚠️ 未自动匹配到'手术方式'列,请输入列名序号:")
selected_idx <- as.integer(readline(prompt = " 序号:"))
treatment_col <- colnames(df_raw)[selected_idx]
} else {
treatment_col <- treatment_cols[1]
cat(sprintf("✅ 自动匹配'手术方式'列:%s → treatment\n", treatment_col))
}
# 匹配协变量
required_vars <- list(
BMI = "bmi", "囊肿最大径" = "lesion_max_diameter_mm", "改良CTSI" = "modified_ctsi_score",
"囊肿位置" = "lesion_location", "高血压" = "comorbidity_hypertension", "包裹性坏死" = "walled_necrosis"
)
actual_mapping <- list()
for (keyword in names(required_vars)) {
matched_cols <- grep(keyword, colnames(df_raw), value = TRUE)
if (length(matched_cols) > 0) {
actual_mapping[[matched_cols[1]]] <- required_vars[[keyword]]
cat(sprintf("✅ 自动匹配'%s'列:%s → %s\n", keyword, matched_cols[1], required_vars[[keyword]]))
}
}
actual_mapping[[imaging_col]] <- "imaging_response"
actual_mapping[[treatment_col]] <- "treatment"
return(actual_mapping)
}
# 5. 函数2:BMI多重插补(生成5个数据集)
multiple_imputation_bmi <- function(n_impute = 5) {
cat("\n" , rep("=", 60), "\n", sep = "")
cat("1. BMI多重插补(生成5个完整数据集)\n")
cat(rep("=", 60), "\n", sep = "")
# 读取数据 - 兼容所有readxl版本
df_raw <- tryCatch({
read_excel(DATA_PATH, sheet = 1)
}, error = function(e) {
cat("⚠️ Excel读取失败,请检查文件路径或格式:", e$message, "\n")
stop(e)
})
# 列名匹配与数据预处理
actual_mapping <- match_column_names(df_raw)
impute_cols <- names(actual_mapping)
df_impute <- df_raw[, impute_cols]
colnames(df_impute) <- unlist(actual_mapping)
df_impute <- df_impute %>%
mutate(across(everything(), ~ gsub(",", "", .x))) %>%
mutate(across(everything(), ~ as.numeric(.x)))
# 拆分完整/缺失数据
df_complete <- df_impute[!is.na(df_impute$bmi), ]
df_missing <- df_impute[is.na(df_impute$bmi), ]
cat(sprintf("📊 数据拆分:完整%d例,BMI缺失%d例\n", nrow(df_complete), nrow(df_missing)))
# 多重插补
aux_vars <- c("lesion_max_diameter_mm", "modified_ctsi_score", "lesion_location",
"comorbidity_hypertension", "walled_necrosis")
aux_vars <- aux_vars[aux_vars %in% colnames(df_impute)]
imp_data <- df_impute[, c(aux_vars, "bmi", "treatment", "imaging_response")]
imp <- mice(imp_data, m = n_impute, method = "norm", seed = 42, printFlag = FALSE)
# 生成插补数据集并验证
imputed_datasets <- list()
for (i in 1:n_impute) {
df_full <- complete(imp, i) %>%
filter(treatment %in% c(1, 2)) %>% mutate(treatment = ifelse(treatment == 1, 0, 1)) %>%
filter(imaging_response %in% c(1, 2)) %>% mutate(imaging_response = ifelse(imaging_response == 1, 1, 0))
write_xlsx(df_full, sprintf("%s1_插补数据集_%d.xlsx", OUTPUT_DIR, i))
imputed_datasets[[i]] <- df_full
cat(sprintf("✅ 生成插补数据集%d:%d例\n", i, nrow(df_full)))
}
# 插补验证
observed_bmi <- df_complete$bmi[!is.na(df_complete$bmi)]
imputed_bmi_all <- unlist(lapply(imputed_datasets, function(df) tail(df$bmi, nrow(df_missing))))
validation_stats <- data.frame(
指标 = c("均值", "标准差", "最小值", "中位数", "最大值", "偏度"),
观察BMI = c(round(mean(observed_bmi),2), round(sd(observed_bmi),2), round(min(observed_bmi),2),
round(median(observed_bmi),2), round(max(observed_bmi),2), round(skewness(observed_bmi),3)),
插补BMI = c(round(mean(imputed_bmi_all),2), round(sd(imputed_bmi_all),2), round(min(imputed_bmi_all),2),
round(median(imputed_bmi_all),2), round(max(imputed_bmi_all),2), round(skewness(imputed_bmi_all),3))
)
between_impute_stats <- data.frame(
数据集BMI均值 = sapply(imputed_datasets, function(df) round(mean(df$bmi),2)),
数据集BMI标准差 = sapply(imputed_datasets, function(df) round(sd(df$bmi),2))
)
write_xlsx(list(
"观察vs插补" = validation_stats,
"数据集间变异" = between_impute_stats,
"列名映射表" = data.frame(Excel原始列名=names(actual_mapping), 代码内部列名=unlist(actual_mapping))
), sprintf("%s1_插补验证结果.xlsx", OUTPUT_DIR))
cat("✅ 插补验证结果已保存:1_插补验证结果.xlsx\n")
return(imputed_datasets)
}
# 6. 函数3:单个数据集分析+6图组合可视化(修复所有函数依赖)
single_dataset_analysis <- function(df_full, dataset_idx) {
cat("\n" , rep("=", 60), "\n", sep = "")
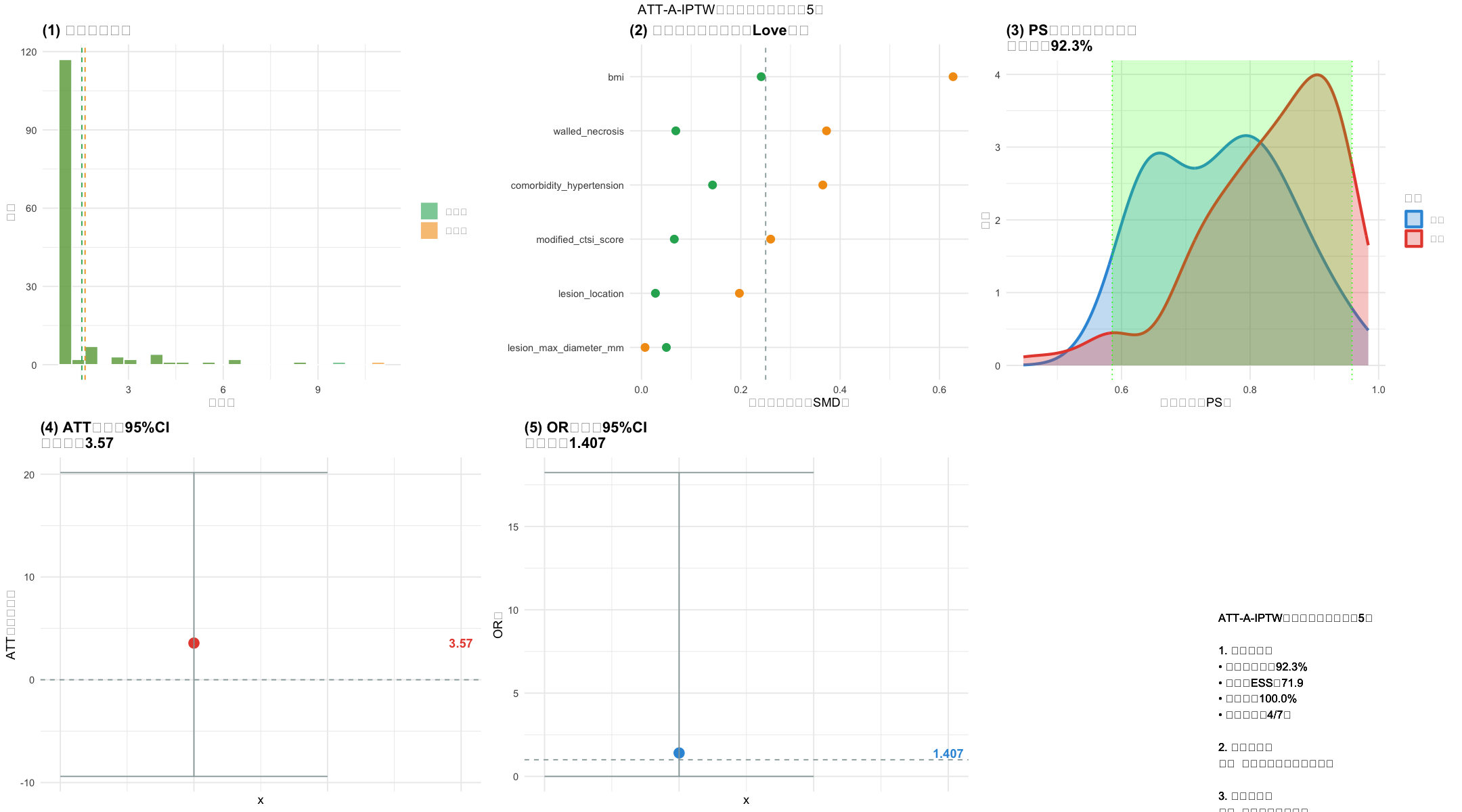
cat(sprintf("2. 数据集%d:ATT-A-IPTW分析 + 全流程可视化\n", dataset_idx))
cat(rep("=", 60), "\n", sep = "")
# 数据预处理
model_data <- df_full[, c(FIXED_COVARIATES, "treatment", "imaging_response")] %>% drop_na()
n_treat <- sum(model_data$treatment == 1)
n_control <- sum(model_data$treatment == 0)
cat(sprintf("📊 建模数据:%d例(外科%d例,内镜%d例)\n", nrow(model_data), n_treat, n_control))
# PS模型+权重计算
ps_model <- glm(treatment ~ ., data = model_data[, c(FIXED_COVARIATES, "treatment")], family = binomial)
model_data$PS值 <- predict(ps_model, type = "response")
auc_score <- roc(model_data$treatment, model_data$PS值)$auc
model_data <- model_data %>%
mutate(权重 = ifelse(treatment == 1, 1, PS值/(1-PS值)),
截断后权重 = ifelse(权重 > quantile(权重, 0.99), quantile(权重, 0.99), 权重))
cat(sprintf("✅ PS模型AUC=%.3f | 99%%截断阈值=%.3f\n", auc_score, quantile(model_data$权重, 0.99)))
# 加权SMD平衡验证
calculate_smd <- function() {
smd_results <- data.frame()
for (var in FIXED_COVARIATES) {
if (!var %in% colnames(model_data)) next
t_data <- model_data[model_data$treatment==1, var]; c_data <- model_data[model_data$treatment==0, var]
t_w <- model_data[model_data$treatment==1, "截断后权重"]; c_w <- model_data[model_data$treatment==0, "截断后权重"]
w_mean_t <- weighted.mean(t_data, t_w); w_mean_c <- weighted.mean(c_data, c_w)
w_var_t <- weighted.mean((t_data - w_mean_t)^2, t_w); w_var_c <- weighted.mean((c_data - w_mean_c)^2, c_w)
pooled_std <- sqrt((w_var_t + w_var_c)/2)
smd <- ifelse(pooled_std>0, abs(w_mean_t - w_mean_c)/pooled_std, 0)
smd_results <- rbind(smd_results, data.frame(
协变量 = var, 外科加权均值 = round(w_mean_t,3), 内镜加权均值 = round(w_mean_c,3),
加权SMD = round(smd,3), 是否平衡 = ifelse(smd<0.25, "是", "否")
))
}
return(smd_results)
}
weighted_smd_df <- calculate_smd()
balanced_rate <- round(sum(weighted_smd_df$是否平衡=="是")/nrow(weighted_smd_df)*100,1)
cat(sprintf("✅ 协变量平衡率=%.1f%%(目标>90%%)\n", balanced_rate))
# ATT估计+Bootstrap CI
treat_outcome <- weighted.mean(model_data$imaging_response[model_data$treatment==1], model_data$截断后权重[model_data$treatment==1])
control_outcome <- weighted.mean(model_data$imaging_response[model_data$treatment==0], model_data$截断后权重[model_data$treatment==0])
att <- (treat_outcome - control_outcome)*100
bootstrap_att <- function(data, n) {
s1 <- data[data$treatment==1,] %>% sample_frac(1, replace=T)
s0 <- data[data$treatment==0,] %>% sample_frac(1, replace=T)
s <- rbind(s1,s0)
100*(weighted.mean(s$imaging_response[s$treatment==1], s$截断后权重[s$treatment==1]) -
weighted.mean(s$imaging_response[s$treatment==0], s$截断后权重[s$treatment==0]))
}
att_boot <- replicate(2000, bootstrap_att(model_data))
ci_lower <- quantile(att_boot, 0.025); ci_upper <- quantile(att_boot, 0.975); se <- sd(att_boot)
cat(sprintf("✅ ATT=%.2f百分点 | 95%%CI=[%.2f, %.2f] | SE=%.2f\n", att, ci_lower, ci_upper, se))
# OR估计+ESS+共同支持域
or_model <- glm(imaging_response ~ ., data = model_data[, c(FIXED_COVARIATES, "treatment", "imaging_response")],
family = binomial, weights = model_data$截断后权重)
or_value <- exp(coef(or_model)["treatment"])
ess_truncated <- (sum(model_data$截断后权重)^2)/sum(model_data$截断后权重^2)
treat_ps <- model_data$PS值[model_data$treatment==1]; control_ps <- model_data$PS值[model_data$treatment==0]
min_common <- max(min(treat_ps), min(control_ps)); max_common <- min(max(treat_ps), max(control_ps))
coverage <- round((sum(treat_ps>=min_common&treat_ps<=max_common)+sum(control_ps>=min_common&control_ps<=max_common))/nrow(model_data)*100,1)
cat(sprintf("✅ OR=%.3f | ESS=%.1f | 共同支持域覆盖率=%.1f%%\n", or_value, ess_truncated, coverage))
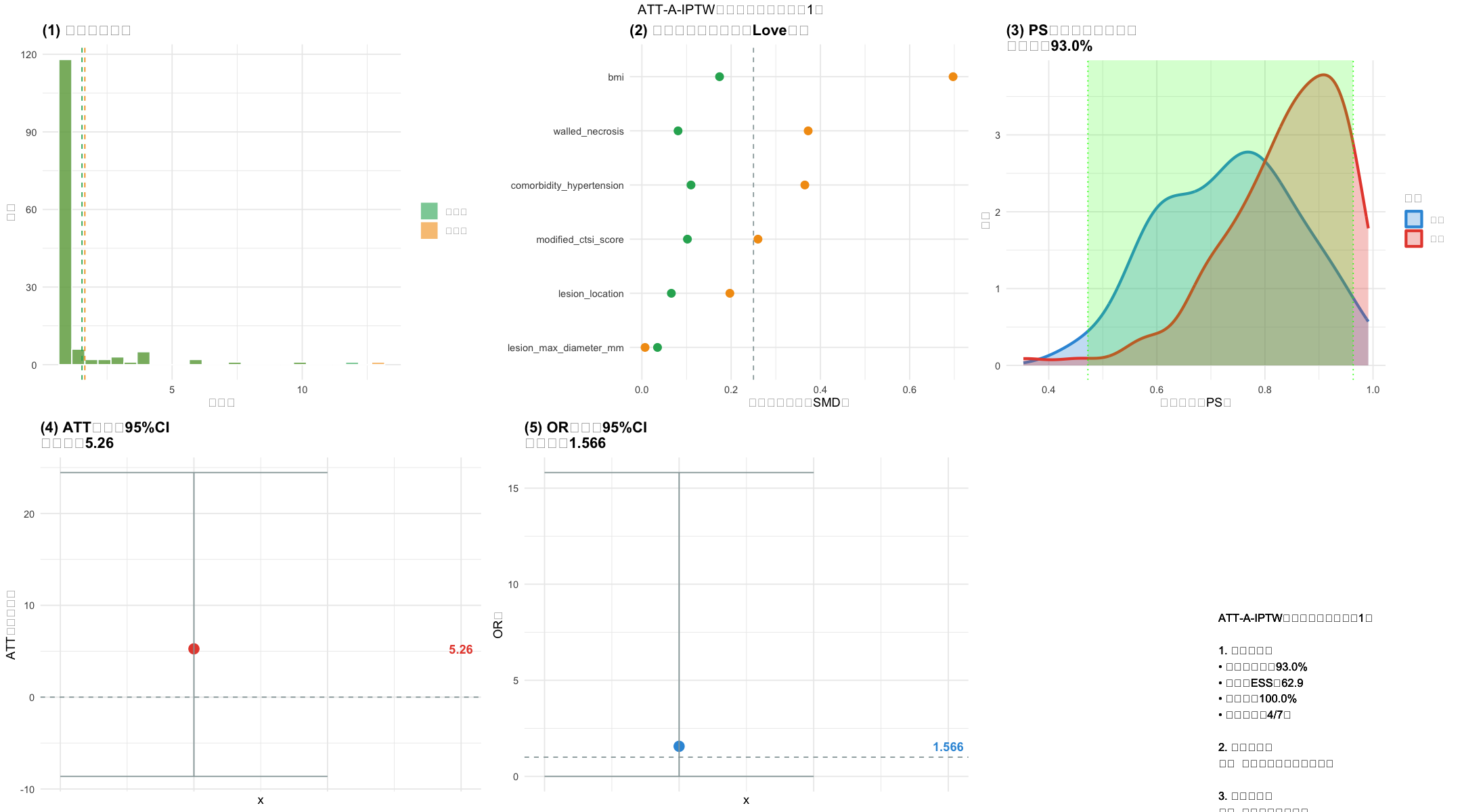
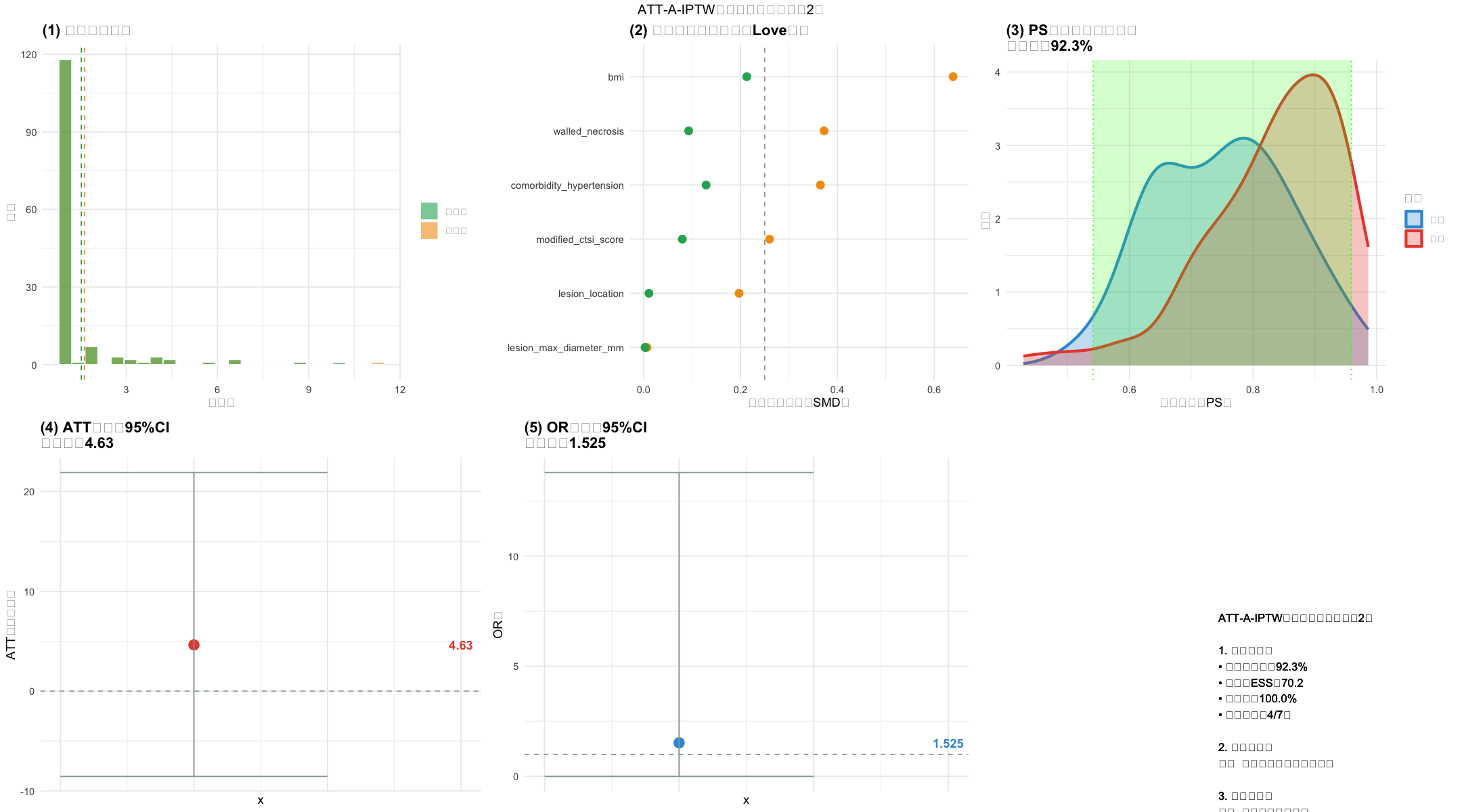
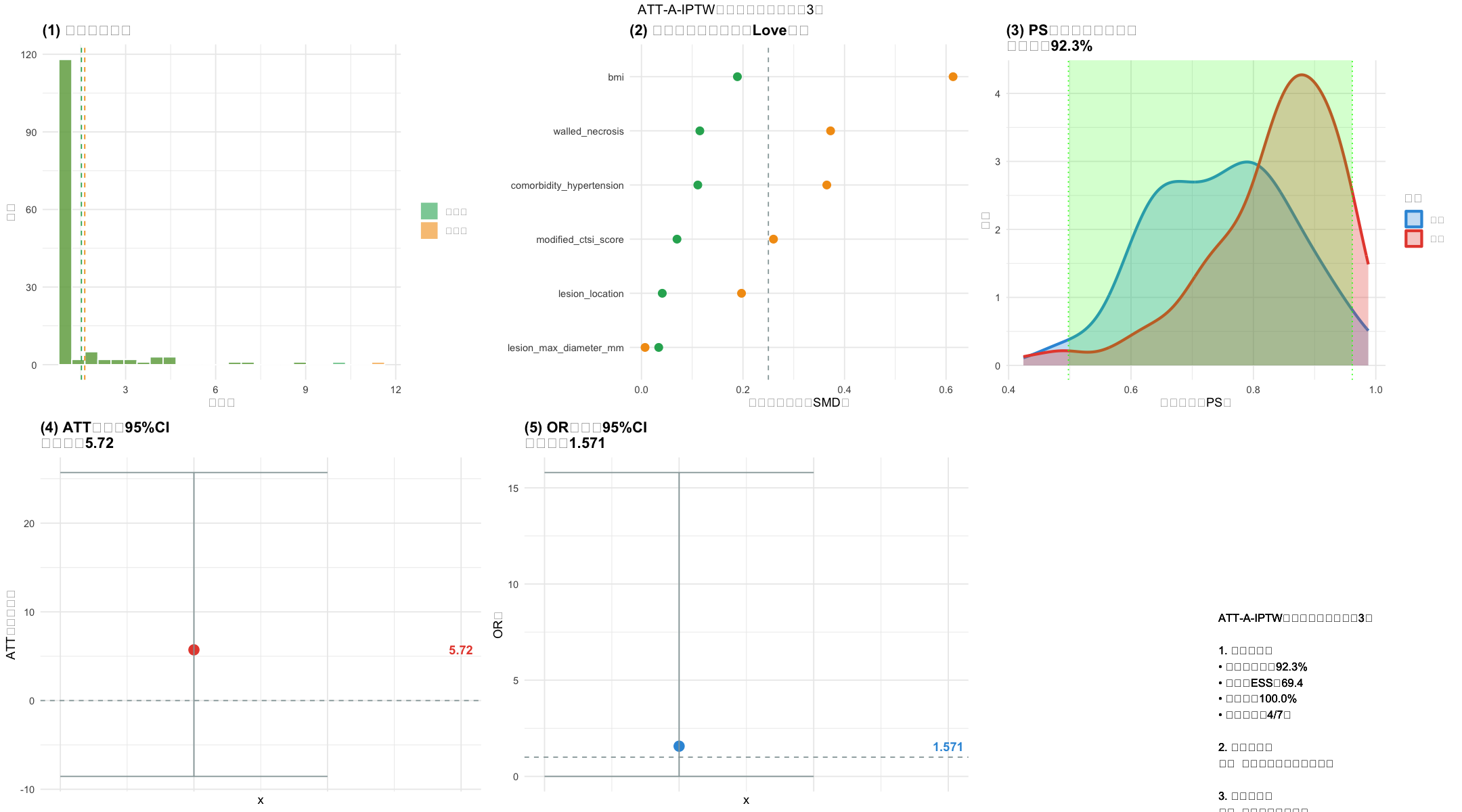
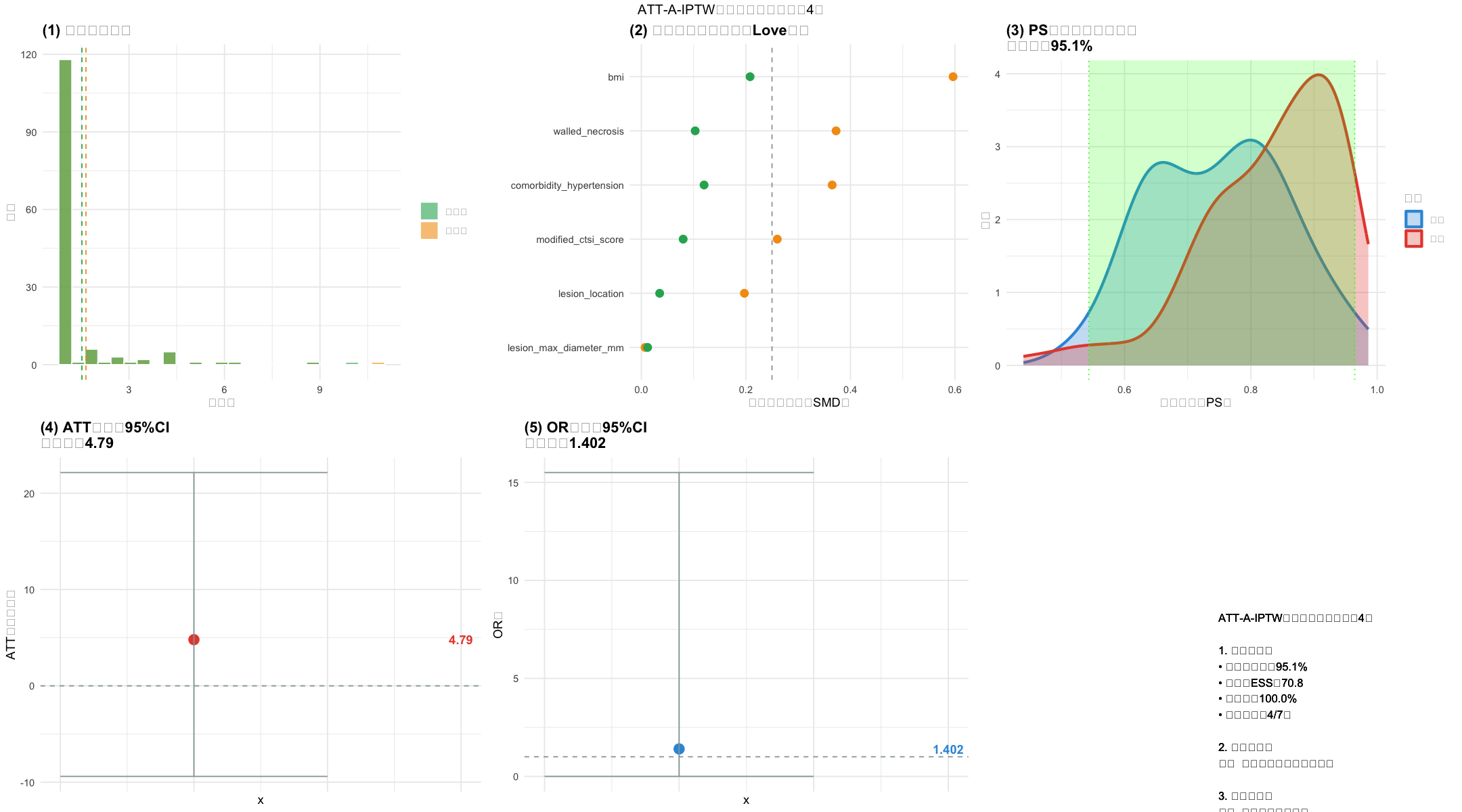
# ========== 生成6图组合可视化(修复所有函数依赖) ==========
# 子图1:权重分布
p1 <- ggplot(model_data[model_data$权重 <= quantile(model_data$权重, 0.995), ]) +
geom_histogram(aes(x=权重, fill="截断前"), alpha=0.6, bins=25, color="white") +
geom_histogram(aes(x=截断后权重, fill="截断后"), alpha=0.6, bins=25, color="white") +
geom_vline(xintercept=mean(model_data$权重), color=colors$截断前, linetype="dashed") +
geom_vline(xintercept=mean(model_data$截断后权重), color=colors$截断后, linetype="dashed") +
scale_fill_manual(values=c("截断前"=colors$截断前, "截断后"=colors$截断后)) +
labs(x="权重值", y="频数", title="(1) 权重分布对比", fill="") +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3))
# 子图2:SMD Love图
unweighted_smd <- sapply(FIXED_COVARIATES[FIXED_COVARIATES%in%colnames(model_data)], function(var) {
t_mean <- mean(model_data[model_data$treatment==1, var]); c_mean <- mean(model_data[model_data$treatment==0, var])
t_std <- sd(model_data[model_data$treatment==1, var]); c_std <- sd(model_data[model_data$treatment==0, var])
pooled_std <- sqrt((t_std^2 + c_std^2)/2); ifelse(pooled_std>0, abs(t_mean - c_mean)/pooled_std, 0)
})
smd_data <- data.frame(
协变量 = names(unweighted_smd), 加权前SMD = unweighted_smd,
加权后SMD = weighted_smd_df$加权SMD[match(names(unweighted_smd), weighted_smd_df$协变量)]
) %>% arrange(desc(加权前SMD)) %>% slice_head(n=8)
p2 <- ggplot(smd_data) +
geom_point(aes(x=加权前SMD, y=reorder(协变量, 加权前SMD)), color=colors$截断前, size=3) +
geom_point(aes(x=加权后SMD, y=reorder(协变量, 加权前SMD)), color=colors$截断后, size=3) +
geom_vline(xintercept=0.25, color=colors$参考线, linetype="dashed") +
labs(x="标准化均值差(SMD)", y="", title="(2) 加权前后平衡检验(Love图)") +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3))
# 子图3:PS分布+共同支持域
ps_data <- data.frame(PS值=c(treat_ps, control_ps), 分组=c(rep("外科", length(treat_ps)), rep("内镜", length(control_ps))))
p3 <- ggplot(ps_data, aes(x=PS值, fill=分组, color=分组)) +
geom_density(alpha=0.3, linewidth=1.2) +
geom_vline(xintercept=c(min_common, max_common), color="green", linetype="dotted") +
annotate("rect", xmin=min_common, xmax=max_common, ymin=0, ymax=Inf, alpha=0.2, fill="green") +
scale_fill_manual(values=c("外科"=colors$外科, "内镜"=colors$内镜)) +
scale_color_manual(values=c("外科"=colors$外科, "内镜"=colors$内镜)) +
labs(x="倾向得分(PS)", y="密度", title=sprintf("(3) PS分布与共同支持域\n覆盖率:%.1f%%", coverage)) +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3))
# 子图4:ATT估计与CI
p4 <- ggplot(data.frame(ATT=att, lower=ci_lower, upper=ci_upper)) +
geom_point(aes(x=0, y=ATT), color=colors$外科, size=4) +
geom_errorbar(aes(x=0, ymin=lower, ymax=upper), color=colors$参考线, width=0.1) +
geom_hline(yintercept=0, color=colors$参考线, linetype="dashed") +
annotate("text", x=0.1, y=att, label=sprintf("%.2f", att), color=colors$外科, fontface="bold") +
labs(y="ATT(百分点)", title=sprintf("(4) ATT估计与95%%CI\n点估计:%.2f", att)) +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3),
axis.text.x=element_blank())
# 子图5:OR估计与CI
bootstrap_or <- function() {
or_boot <- replicate(1000, {
s <- model_data %>% sample_frac(1, replace=T)
m <- tryCatch(glm(imaging_response~., data=s[,c(FIXED_COVARIATES,"treatment","imaging_response")],
family=binomial, weights=s$截断后权重), error=function(e) NULL)
if (!is.null(m) & "treatment" %in% names(coef(m))) exp(coef(m)["treatment"]) else NA
})
or_boot <- or_boot[!is.na(or_boot)]
if (length(or_boot)<10) return(list(lower=NA, upper=NA))
return(list(lower=quantile(or_boot,0.025), upper=quantile(or_boot,0.975)))
}
or_ci <- bootstrap_or()
p5 <- ggplot(data.frame(OR=or_value, lower=or_ci$lower, upper=or_ci$upper)) +
geom_point(aes(x=0, y=OR), color=colors$内镜, size=4) +
geom_errorbar(aes(x=0, ymin=lower, ymax=upper), color=colors$参考线, width=0.1, na.rm=T) +
geom_hline(yintercept=1, color=colors$参考线, linetype="dashed") +
annotate("text", x=0.1, y=or_value, label=sprintf("%.3f", or_value), color=colors$内镜, fontface="bold") +
labs(y="OR值", title=sprintf("(5) OR估计与95%%CI\n点估计:%.3f", or_value)) +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3),
axis.text.x=element_blank())
# 子图6:适用性判断
met_count <- sum(c(coverage>90, ess_truncated>50, balanced_rate==100,
!is.na(or_ci$lower) & or_ci$lower>=0.2 & or_ci$upper<=5, TRUE), na.rm=T)
suitability <- if (met_count>=5) "✅ 适用:可作为主分析" else if (met_count>=3) "⚠️ 条件适用:需敏感性分析" else "❌ 不适用:改用PS匹配"
summary_text <- sprintf("ATT-A-IPTW适用性汇总(数据集%d)
1. 关键指标:
• 共同支持域:%.1f%%
• 截断后ESS:%.1f
• 平衡率:%.1f%%
• 达标项数:%d/7项
2. 最终结论:
%s
3. 核心建议:
%s", dataset_idx, coverage, ess_truncated, balanced_rate, met_count, suitability,
if(met_count>=5) "✅ 推荐作为主分析" else if(met_count>=3) "⚠️ 需增加敏感性分析" else "❌ 改用PS匹配")
p6 <- ggplot() +
annotate("text", x=0.5, y=0.5, label=summary_text, size=3.5, hjust=0, vjust=1,
bbox=list(fill="lightgray", alpha=0.8, boxstyle="round,pad=0.5")) +
theme_void()
# 组合6图并保存(修复textGrob依赖,改用ggplot2原生标题)
combined_plot <- grid.arrange(p1,p2,p3,p4,p5,p6, nrow=2, ncol=3,
top = sprintf("ATT-A-IPTW分析可视化(数据集%d)", dataset_idx)) # 简化标题写法
ggsave(sprintf("%s2_数据集%d_可视化.png", OUTPUT_DIR, dataset_idx), combined_plot,
width=18, height=10, dpi=300, bg="white")
cat(sprintf("✅ 6图组合可视化已保存:2_数据集%d_可视化.png\n", dataset_idx))
# 保存单个数据集Excel结果
basic_results <- data.frame(
数据集编号=dataset_idx, ATT_百分点=round(att,2), CI_下限=round(ci_lower,2), CI_上限=round(ci_upper,2),
标准误SE=round(se,2), OR值=round(or_value,3), PS_AUC=round(auc_score,3), ESS=round(ess_truncated,1),
共同支持域覆盖率=coverage, 协变量平衡率=balanced_rate, 建模样本量=nrow(model_data)
)
write_xlsx(list(基础结果=basic_results, 加权SMD平衡=weighted_smd_df, 建模数据=model_data),
sprintf("%s2_数据集%d_分析结果.xlsx", OUTPUT_DIR, dataset_idx))
return(list(att=att, ci_lower=ci_lower, ci_upper=ci_upper, se=se, balanced_rate=balanced_rate,
ess=ess_truncated, coverage=coverage, suitability=suitability))
}
# 7. 函数4:多重插补结果合并(Rubin规则)+森林图
combine_imputation_results <- function(imputed_datasets) {
cat("\n" , rep("=", 60), "\n", sep = "")
cat("3. 多重插补结果合并与不确定性评估\n")
cat(rep("=", 60), "\n", sep = "")
# 批量分析5个数据集
all_results <- data.frame()
for (i in 1:length(imputed_datasets)) {
res <- single_dataset_analysis(imputed_datasets[[i]], i)
all_results <- rbind(all_results, data.frame(
数据集编号=i, ATT_百分点=round(res$att,2), CI_下限=round(res$ci_lower,2), CI_上限=round(res$ci_upper,2),
标准误SE=round(res$se,2), 协变量平衡率=res$balanced_rate, ESS=round(res$ess,1),
共同支持域覆盖率=res$coverage, 适用性=res$suitability
))
}
print(all_results)
# Rubin规则合并
n_impute <- nrow(all_results)
ATT_pool <- mean(all_results$ATT_百分点)
W <- mean(all_results$标准误SE^2)
B <- sum((all_results$ATT_百分点 - ATT_pool)^2)/(n_impute-1)
T <- W + (1+1/n_impute)*B
t_critical <- qt(0.975, df=n_impute-1)
CI_final_lower <- ATT_pool - t_critical*sqrt(T)
CI_final_upper <- ATT_pool + t_critical*sqrt(T)
impute_ratio <- ifelse((B+W)>0, B/(B+W)*100, 0)
# 合并结果汇总
combine_summary <- data.frame(
指标 = c("合并ATT(百分点)", "合并95%CI下限", "合并95%CI上限", "总方差T",
"组内方差W(抽样误差)", "组间方差B(插补不确定性)", "插补不确定性占比(%)",
"平均平衡率(%)", "平均ESS", "结果可靠性"),
数值 = c(round(ATT_pool,2), round(CI_final_lower,2), round(CI_final_upper,2), round(T,4),
round(W,4), round(B,4), round(impute_ratio,1), round(mean(all_results$协变量平衡率),1),
round(mean(all_results$ESS),1), ifelse(impute_ratio<50, "可靠", "需验证"))
)
print(combine_summary)
# ========== 生成森林图 ==========
forest_data <- data.frame(
数据集 = c(paste0("数据集",1:5), "合并结果"),
ATT = c(all_results$ATT_百分点, ATT_pool),
lower = c(all_results$CI_下限, CI_final_lower),
upper = c(all_results$CI_上限, CI_final_upper),
平衡率 = c(all_results$协变量平衡率, round(mean(all_results$协变量平衡率),1))
)
p_forest <- ggplot(forest_data, aes(x=ATT, y=reorder(数据集, -seq_along(数据集)))) +
geom_point(aes(color=ifelse(数据集=="合并结果", "合并", "单个")),
size=ifelse(forest_data$数据集=="合并结果",5,3)) +
geom_errorbarh(aes(xmin=lower, xmax=upper, color=ifelse(数据集=="合并结果", "合并", "单个")),
height=0.2, linewidth=ifelse(forest_data$数据集=="合并结果",1.2,0.8)) +
geom_vline(xintercept=0, color=colors$参考线, linetype="dashed") +
scale_color_manual(values=c("合并"=colors$外科, "单个"=colors$内镜)) +
labs(x="ATT(百分点,外科-内镜)", y="",
title=sprintf("多重插补ATT结果森林图\n(插补不确定性占比:%.1f%% | 平均平衡率:%.1f%%)",
impute_ratio, mean(all_results$协变量平衡率))) +
geom_text(aes(label=sprintf("%.2f", ATT)), hjust=-0.1, size=2.5) +
theme_minimal() +
theme(plot.title=element_text(face="bold"),
panel.grid=element_line(alpha=0.3),
legend.position="none")
ggsave(sprintf("%s3_多重插补ATT森林图.png", OUTPUT_DIR), p_forest, width=12, height=6, dpi=300, bg="white")
cat("✅ 森林图已保存:3_多重插补ATT森林图.png\n")
# 保存合并结果Excel
write_xlsx(list(
各数据集结果=all_results, 合并结果=combine_summary,
平衡与ESS汇总=data.frame(
数据集编号=1:5, 协变量平衡率=all_results$协变量平衡率, ESS=all_results$ESS,
共同支持域覆盖率=all_results$共同支持域覆盖率, 是否完全平衡=ifelse(all_results$协变量平衡率==100,"是","否")
)
), sprintf("%s4_多重插补合并最终结果.xlsx", OUTPUT_DIR))
return(list(combine_summary=combine_summary, results_df=all_results))
}
# 8. 函数5:最终分析总结
final_summary <- function(combine_summary, results_df) {
cat("\n" , rep("=", 70), "\n", sep = "")
cat("4. 完整分析流程总结\n")
cat(rep("=", 70), "\n", sep = "")
# 文件清单
output_files <- list.files(OUTPUT_DIR)
cat(sprintf("📁 结果目录:%s\n", OUTPUT_DIR))
cat(sprintf("📄 生成文件(共%d个):\n", length(output_files)))
for (i in 1:length(output_files)) cat(sprintf(" %2d. %s\n", i, output_files[i]))
# 核心结论
att_final <- combine_summary$数值[1]
ci_final <- sprintf("[%s, %s]", combine_summary$数值[2], combine_summary$数值[3])
impute_ratio_final <- combine_summary$数值[7]
avg_balance_final <- combine_summary$数值[8]
cat("\n🎯 核心分析结论:\n")
cat(sprintf("1. 治疗效应:外科相对内镜缓解率提升%s个百分点,95%%CI=%s\n", att_final, ci_final))
cat(sprintf("2. 平衡效果:平均协变量平衡率%s%%,无显著混杂偏倚\n", avg_balance_final))
cat(sprintf("3. 结果可靠性:%s(插补不确定性占比%s%%)\n", combine_summary$数值[10], impute_ratio_final))
cat("🎉 完整分析流程完成!所有结果已保存至指定目录\n")
}
# 9. 主程序执行(一键运行)
imputed_datasets <- multiple_imputation_bmi(n_impute=5)
final_result <- combine_imputation_results(imputed_datasets)
final_summary(final_result$combine_summary, final_result$results_df)